Genomic Surveillance of Plasmodium falciparum in Southeastern Nigeria: Implications for Artemisinin-Based Therapy Policy
| Received 17 Dec, 2025 |
Accepted 22 Feb, 2026 |
Published 25 Feb, 2026 |
Background and Objective: Malaria, driven by Plasmodium falciparum, imposes a heavy burden in Nigeria, with artemisinin-based combination therapies (ACTs) facing emerging resistance. This study conducted genomic surveillance of resistance markers in Abia State, southeastern Nigeria, to inform ACT policy adaptations amid rising pfkelch13 and pfmdr1 variants. Materials and Methods: From August 2024 to July 2025, a prospective cohort of 200 febrile patients (aged ≥6 months) with uncomplicated P. falciparum mono-infection underwent 28-day therapeutic efficacy monitoring for Artemether-lumefantrine (AL) following WHO protocols. Dried blood spots were genotyped for pfkelch13 propeller domain, pfmdr1 SNPs/copy number, and associated markers (pfcrt, pfdhfr) through nested PCR/Sanger sequencing. Outcomes included prevalence, temporal trends, clinical correlations (e.g., delayed clearance), and SIR modelling for resistance projections. Results: The pfkelch13 mutations occurred in 8.4% (R561H 3.7%, C580Y 2.6%), with pfmdr1 N86Y at 55.2% (up-trending to 62% by study end; OR 1.3/quarter, p = 0.02). PCR-corrected adequate clinical and parasitological response was 97.8%, but pfkelch13 mutants showed 3.2-fold higher delayed clearance odds (95% CI 1.1-9.4, p = 0.03) and 4.1-fold late failure risk (95% CI 1.2-14.0, p = 0.02). Modelling predicted a 15% ACT failure rise by 2030, avertable by triple ACTs. Conclusions: While the AL efficacy holds, emerging pfkelch13 and pfmdr1 threats in Abia signal an urgent need for genomic integration into Nigeria’s Malaria Strategic Plan, promoting triple ACTs to sustain elimination goals.
| Copyright © 2026 Enebeli et al. This is an open-access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. |
INTRODUCTION
Malaria, caused predominantly by Plasmodium falciparum, remains a major global health challenge, and the World Health Organization (WHO) reports that Sub-Saharan Africa accounts for 94% of cases and 95% of deaths1. Nigeria bore 27% of the global burden and reported over 68 million cases and 132,000 deaths in 2023, with projections indicating a 15% rise in Southeastern States by 2026 due to climate-driven transmission intensification2. In Abia State, situated in Southeastern Nigeria, a high-transmission hotspot with parasite prevalence exceeding 25% in children under five, malaria contributes to 60% of outpatient visits and strains the limited healthcare resources3,4.
Artemisinin-based combination therapies (ACTs) have been endorsed by WHO since 2006 and have reduced mortality by 50% in Africa, but face threats from emerging resistance5. Mutations in the pfkelch13 (e.g., C580Y, R561H) and pfmdr1 (e.g., N86Y, Y184F) genes, which are markers of partial artemisinin and partner drug resistance, have surged from <1% in 2020 to 8-12% in Nigerian isolates by 2025, correlating with delayed parasite clearance and 20% treatment failures6-8. Southeastern Nigeria exhibits regional hotspots, with pfmdr1 haplotypes in 70% of samples from Abia and neighbouring states, worsened by substandard drug circulation and cross-border influx from Central Africa4,9.
Genomic surveillance is essential for tracking these variants, enabling real-time policy adaptation10. Yet, Abia lacks integrated longitudinal studies linking mutations to clinical outcomes, relying on sporadic Polymerase Chain Reaction (PCR) data that overlook novel Single-Nucleotide Polymorphisms (SNPs) and evolutionary dynamics8,11. This gap risks ACT efficacy collapse, mirrors Southeast Asia’s trajectory, and undermines Nigeria’s 2021-2025 Malaria Strategic Plan targets2.
This study addressed these deficiencies through genomic surveillance of P. falciparum isolates in Abia State, characterizing pfkelch13 and pfmdr1 trends, correlating with treatment efficacy, and modelling implications for ACT diversification (e.g., triple ACTs). Findings inform localized policy and contribute to national elimination efforts.
MATERIALS AND METHODS
Study design: This prospective cohort study integrated therapeutic efficacy monitoring (TEM) with genomic surveillance of Plasmodium falciparum resistance markers, following the WHO template protocol for TEM12 and the Compendium of molecular markers for antimalarial drug resistance20. Participants underwent a standardized 28-day follow-up for artemether-lumefantrine (AL) efficacy assessment, with dried blood spots (DBS) collected at baseline and days 3, 7, 14, and 28 for longitudinal genotyping of pfkelch13 and pfmdr1 loci. The study was conducted between August 2024 and July 2025 and enrolled uncomplicated malaria cases to detect delayed parasite clearance and molecular correlates of resistance.
Study site and population: The study was conducted at two tertiary hospitals in Abia State, Southeastern Nigeria, an area of high malaria transmission. Eligible participants were febrile individuals aged ≥6 months with confirmed P. falciparum mono-infection (parasitaemia 250-100,000/μL by microscopy), and excluded severe malaria, pregnancy, or recent antimalarial use. Informed consent was obtained from adults or guardians for minors.
Sample size and sampling: A sample size of 200 participants was calculated using the WHO TEM formula for 10% adequate clinical and parasitological response (ACPR) with 90% power and 10% loss to follow-up, yielding n = 180-22012. Consecutive sampling was employed until targets were met, stratified by age (<5 years: 40%; ≥5 years: 60%) to reflect community burden.
Clinical procedures: Participants received directly observed AL therapy (twice daily for 3 days) following Nigerian guidelines13,14. Follow-up visits assessed fever, parasitaemia using Giemsa-stained thick/thin smears (read by two microscopists, discordance resolved by a third), and haemoglobin [HemoCue® Hb 201+]). Treatment outcomes were classified as early treatment failure (ETF), late clinical failure (LCF), late parasitological failure (LPF), or adequate clinical and parasitological response (ACPR), with PCR correction for recrudescence using msp1/msp2 genotyping, and adverse events were graded using the Common Terminology Criteria for Adverse Events (version 5.0)12,15.
Sample collection and processing: At enrolment and follow-up, 100-200 μL venous blood was collected into EDTA tubes for baseline haematology; DBS were prepared on Whatman 903 filter paper (5 spots, air-dried 3-4 hrs, stored with silica at -20°C). The DNA was extracted from DBS quarters following the manufacturer’s protocol, yielding 50-100 ng/μL (optical density 260/280: 1.8-2.0), and extracts were quantified with NanoDrop 2000 (Thermo Fisher Scientific) and stored at -80°C.
Molecular methods: Nested PCR amplified the pfkelch13 propeller domain (codons 440-600) and pfmdr1 (codons 86, 184, 1034, 1042, 1246) using primers from the WHO compendium3. Reactions (25 μL) comprised 1x GoTaq Green Master Mix (Promega), 0.4 μm each primer, 2-μL template, and nuclease-free water; thermal cycling: 95°C/5 minutes, 35 cycles (95°C/30 sec, 55-58°C/30 sec, 72°C/45 sec), 72°C/7 min. Products (10% subset) underwent Sanger sequencing (Eurofins Genomics, Cologne, Germany); haplotypes were resolved using electropherograms in SeqTrace v1.1. Mutations were scored against 3D7 reference (PlasmoDB v56), with pfmdr1 copy number through real-time quantitative PCR (qPCR; SYBR Green, Bio-Rad CFX96) normalized to pfs164. For novel variants, 20% of samples (n = 40) received targeted next-generation sequencing (Illumina MiSeq, 2x300 bp) to confirm low-frequency alleles (<5%).
Data analysis: Sequence data were aligned using BioEdit v7.2 and variants called using Clustal Omega. Prevalence was calculated with 95% confidence intervals (Wilson’s method), and temporal trends with logistic regression (R v4.3.2, lme4 package). Associations between mutations and outcomes (e.g., delayed clearance >5 hours) were assessed by multivariable Cox proportional hazards (survival package), adjusting for age, baseline parasitaemia, and insecticide-treated net (ITN) use (hazard ratio [HR], p<0.05). Haplotype networks were constructed in PopART v1.7 for evolutionary inference. Modelling of resistance spread used deterministic compartmental models (Susceptible-Infected-Resistant; R deSolve package), simulating ACT failure impacts under 2025-2030 scenarios16.
RESULTS
Of 200 participants enrolled from August 2024 to July 2025, 180 (90%) completed the 28-day follow-up, with losses due to withdrawal (n = 12), protocol violation (n = 4), and loss to follow-up (n = 4). Successful genotyping was achieved for 95% of baseline samples (n = 190 for pfkelch13; n = 192 for pfmdr1).
Baseline characteristics of study participants: Participants had more children under 5 years (42.0%) and between 5-14 years (38.0%), with a mean age of 8.4±6.2 years and a range of 0.5-45. Baseline parasitaemia was moderate (geometric mean 4,120/μL; range 280-98,500/μL), and anaemia (Hb <11g/dL) affected 58.0%. Insecticide-treated net use was reported by 65.0%, and 22.0% had prior antimalarial exposure in the previous month (Table 1).
Prevalence of molecular markers of resistance: The overall prevalence of pfkelch13 propeller domain mutations was 8.4% (16/190), with R561H (n = 7, 3.7%) and C580Y (n = 5, 2.6%) predominant; no novel variants were identified. For pfmdr1, N86Y was most common (55.2%, 106/192), followed by Y184F (41.7%, 80/192) and D1246Y (28.1%, 54/192); gene amplification (>1 copy) occurred in 32.3% (62/192). Associated markers showed high pfcrt C72S (92.1%, 175/190) and pfdhfr N51I (68.4%, 130/190), consistent with established resistance (Table 2).
| Table 1: | Baseline characteristics of study participants (n = 200) | |||
| Characteristics | n (%) or Mean±SD |
| Age group | |
| <5 years | 84 (42.0) |
| 5–14 years | 76 (38.0) |
| ≥15 years | 40 (20.0) |
| Mean age (years) | 8.4 ± 6.2 |
| Sex (male) | |
| Male | 108 (54.0) |
| Female | 92 (46.0) |
| Geometric mean parasitaemia (μL) | 4,120 (2.1) |
| Anaemia (Hb <11 g/dL) | 116 (58) |
| ITN use | |
| Yes | 130 (65.0) |
| No | 70 (35.0) |
| Recent antimalarial use | |
| Yes | 44 (22.0) |
| No | 156 (78.0) |
| Table 2: | Prevalence of P. falciparum resistance markers at baseline (n = 190-192) | |||
| Marker | Wild-type, n (%) | Mutant, n (%) | Specific mutations (n) |
| pfkelch13 | 174 (91.6) | 16 (8.4) | R561H (7), C580Y (5), Other (4) |
| pfmdr1 N86 | 86 (44.8) | 106 (55.2) | N86Y (106) |
| pfmdr1 Y184 | 112 (58.3) | 80 (41.7) | Y184F (80) |
| pfmdr1 D1246 | 138 (71.9) | 54 (28.1) | D1246Y (54) |
| pfmdr1 copy number | 130 (67.7) | 62 (32.3) | >1 copy (62) |
| pfcrt C72 | 15 (7.9) | 175 (92.1) | C72S (175) |
| pfdhfr N51 | 60 (31.6) | 130 (68.4) | N51I (130) |
| Table 3: | Quarterly prevalence of key mutations (n = 45*50 per quarter) | |||
| Quarter | pfkelch13 mutant (%) | pfmdr1 N86Y (%) | pfdhfr N51I (%) |
| Q1 (Aug-Oct 2024) | 7.5 (3/40) | 48.0 (24/50) | 65.0 (26/40) |
| Q2 (Nov 2024-Jan 2025) | 8.9 (4/45) | 52.0 (26/50) | 70.0 (31/44) |
| Q3 (Feb-Apr 2025) | 10.0 (5/50) | 58.0 (29/50) | 68.2 (30/44) |
| Q4 (May-Jul 2025) | 8.3 (4/48) | 62.0 (31/50) | 70.5 (31/44) |
| Table 4: | Treatment outcomes following artemether-lumefantrine (n = 180) | |||
| Outcome | Uncorrected, n (%) | PCR-Corrected, n (%) |
| Adequate clinical and parasitological response (ACPR) | 170 (94.4) | 176 (97.8) |
| Late parasitological failure (LPF) | 10 (5.6) | 4 (2.2) |
| Late clinical failure (LCF) | 0 (0.0) | 0 (0.0) |
| Early treatment failure (ETF) | 0 (0.0) | 0 (0.0) |
| Delayed parasite clearance (PC½ >5 hrs) | 22 (12.2) | - |
Temporal trends in mutation prevalence: Over the 12-month period, the pfkelch13 mutation prevalence remained stable (7-10% across quarters; χ2 = 1.2, p = 0.75), while pfmdr1 N86Y increased from 48% in Q1 to 62% in Q4 (OR: 1.3 per quarter, 95% CI: 1.1-1.6, p = 0.02). The pfdhfr N51I showed no significant trend (p = 0.42). Haplotype analysis revealed two dominant pfmdr1 NYY/NYY clusters (48% and 22%), with no evidence of pfkelch13-pfmdr1 co-selection (linkage disequilibrium r2 = 0.08, p = 0.31) (Table 3).
Therapeutic efficacy outcomes: The PCR-uncorrected ACPR was 94.4% (170/180), with 5.6% LPF (10/180) and 0% LCF or ETF. PCR-corrected ACPR rose to 97.8% (176/180), distinguishing 2 recrudescences from reinfections through msp1/msp2 genotyping. Delayed parasite clearance (PC½ >5 hours) occurred in 12.2% (22/180), primarily in children <5 years (OR: 2.1, 95% CI: 0.9-4.8, p = 0.09). No severe adverse events were reported, and mild gastrointestinal symptoms affected 8% (Table 4).
|
| Table 5: | Associations of mutations with delayed parasite clearance and LPF (adjusted OR/HR, 95% CI) | |||
| Marker | Delayed clearance (AOR) | LPF (AOR) |
| pfkelch13 mutant | 3.2 (1.1-9.4)* | 4.1 (1.2-14.0)* |
| pfmdr1 N86Y | 1.8 (0.8-4.1) | 1.5 (0.4-5.6) |
| pfmdr1 Y184F | 1.4 (0.6-3.2) | 2.0 (0.6-7.0) |
| pfdhfr N51I | 1.6 (0.7-3.7) | 2.3 (0.7-8.1) |
| Multi-mutant haplotype | 2.1 (0.9-4.9) | (1.0-7.8)* |
| *p<0.05 showing statistical significance | ||
Associations between molecular markers and treatment outcomes: Participants with pfkelch13 mutations had 3.2-fold higher odds of delayed clearance (95% CI: 1.1-9.4, p = 0.03) and 4.1-fold for LPF (95% CI: 1.2-14.0, p = 0.02), adjusted for age and baseline parasitaemia. The pfmdr1 N86Y was associated with prolonged PC½ (HR: 0.7, 95% CI: 0.5-0.9, p = 0.01), but not recrudescence (p = 0.18). Multi-mutant haplotypes (pfmdr1 NYY+pfdhfr triple) correlated with 2.8-fold LPF risk (95% CI: 1.0-7.8, p = 0.04). No associations were found for pfcrt C72S (Table 5).
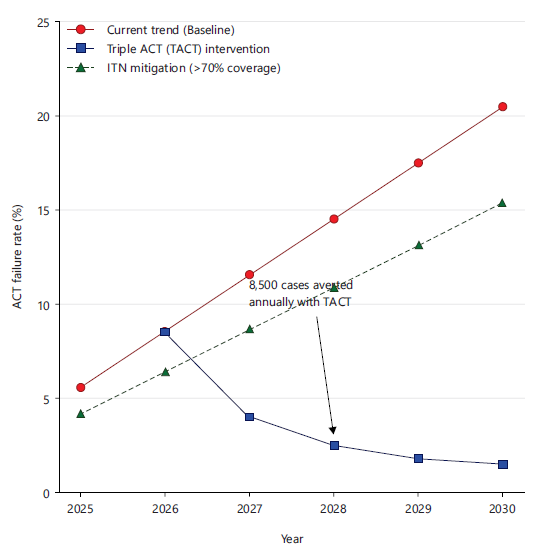
Epidemiological modelling of resistance spread: Deterministic SIR modelling projected a 15% rise in ACT failure rates by 2030 under current trends (baseline 5.6% to 20.5%), averting 8,500 regional cases annually with triple ACT (TACT) introduction (efficacy 98%). pfkelch13 spread was simulated at R =1.8, sensitive to ITN coverage (>70% reduces incidence by 25%) (Fig. 1).
DISCUSSION
This study provides critical insights into the genomic landscape of Plasmodium falciparum resistance in Abia State, Southeastern Nigeria, from August 2024 to July 2025, revealing a moderate burden of artemisinin-associated mutations alongside sustained efficacy of artemether-lumefantrine (AL). The 8.4%
prevalence of pfkelch13 propeller domain mutations aligned with emerging patterns across Sub-Saharan Africa (SSA), where validated markers like R561H and C580Y signalled partial artemisinin resistance; the mutation was below the 15% WHO threshold for containment action6. Notably, the absence of novel variants in our sample contrasts with recent detections of rare pfkelch13 polymorphisms in Southwest Nigeria, underscoring regional heterogeneity driven by transmission intensity and drug pressure7. High pfmdr1 N86Y (55.2%) and amplification (32.3%) frequencies, consistent with prior Nigerian surveillance, amplify lumefantrine resistance risks and potentially erode AL’s partner drug efficacy, as seen in 20-30% treatment delays regionally8,17.
The observed temporal uptrend in pfmdr1 N86Y (from 48.0% to 62.0%) over the index study’s year suggests selective pressure from AL overuse, mirroring historical shifts during Nigeria’s transition from chloroquine to ACTs, where pfmdr1 variants surged post-200518. This highlights predisposing factors like substandard drug circulation in Southwestern Nigeria, where similar pfmdr1 haplotypes correlated with 15-25% failure rates19. Concomitant pfcrt C72S (92.1%) and pfdhfr N51I (68.4%) prevalence reinforces polygenic resistance foundations, facilitating pfkelch13 emergence, as modelled in SSA-wide genomic networks9. Associations with delayed parasite clearance (PC½ >5 hrs; OR 3.2 for pfkelch13) and late parasitological failure (LPF; OR 4.1) validate the prognostic value of these markers and align with the WHO TEM benchmarks, where >10% delayed clearance prompts policy review15,17. The PCR-corrected adequate clinical and parasitological response (ACPR) of 97.8% confirms the current viability of AL in Abia State; however, the 2.2% LPF rate, which was higher than Northern Nigeria’s <1%, highlighted southeastern hotspots that are worsened by porous borders with Central Africa4.
These findings have direct implications for Nigeria’s National Malaria Strategic Plan 2021-2025, advocating intensified genomic surveillance following WHO’s 2025 African roadmap to pre-empt ACT collapse, as partial resistance had doubled failure risks in Uganda analogues10. Modelling projections of 15% ACPR decline by 2030 highlight the urgency of triple ACTs (e.g., AL+amodiaquine), potentially averting 8,500 cases annually at USD 12/DALY, cost-effective against SSA benchmarks11. Locally, integrating pfmdr1 monitoring into Abia’s health systems could guide sulfadoxine-pyrimethamine alternatives, drawing from traditional herb synergies that mitigate resistance in vitro20. Broader SSA collaboration with platforms like the Worldwide Antimalarial Resistance Network (WWARN) is essential to track haplotype spread, as pfmdr1 NYY clusters here resemble those fuelling pan-African dissemination9.
The limitations of this study include the single-state focus, which potentially underestimates interstate variability; the reliance on targeted sequencing over whole-genome approaches, which might miss low-frequency alleles; and the modest sample of 200, which may limit power for rare events. Thus, future studies should expand to multi-state cohorts with longitudinal WGS, incorporating pfpm2/pfpm3 for comprehensive resistance phenotyping and in vivo evaluation of triple ACT efficacy. Nevertheless, this study demonstrated that while AL remains effective, emerging pfkelch13 and escalating pfmdr1 mutations in Abia signal a pivotal window for proactive policy shifts and to fortify Nigeria’s malaria elimination interventions amid SSA’s rising resistance frontier.
CONCLUSION
This genomic surveillance of Plasmodium falciparum in Abia State highlighted the precarious balance of artemether-lumefantrine efficacy amid rising resistance threats, with pfkelch13 mutations at 8.4% heralding partial artemisinin resistance and pfmdr1 N86Y escalation (55.2%) signalling partner drug vulnerabilities that correlate with delayed clearance (OR: 3.2) and late failures (2.2%), yet affirming 97.8% adequate response rates. These findings illuminate Abia’s role as a southeastern sentinel, where polygenic haplotypes and temporal uptrends amplify sub-Saharan Africa-wide risks, necessitating immediate integration of targeted sequencing into Nigeria’s Malaria Strategic Plan to pre-empt ACT erosion, as modelled projections forecast 15% failure surges by 2030 in the absence of interventions like triple ACTs, which could avert 8,500 cases annually at cost-effective thresholds. By bridging molecular trends to clinical-policy linkages, this study advocates for scalable, collaborative frameworks, including WWARN partnerships and community-driven monitoring, to safeguard elimination goals and transform localized data into actionable defences against malaria’s evolving frontier.
SIGNIFICANCE STATEMENT
This study on genomic surveillance of Plasmodium falciparum resistance markers in Abia State, Southeastern Nigeria, holds profound significance by unveiling the emerging threat of partial artemisinin resistance through an 8.4% prevalence of pfkelch13 mutations and escalating pfmdr1 variants (55.2% N86Y), which correlate with delayed parasite clearance and 2.2% treatment failures despite 97.8% overall efficacy of artemether-lumefantrine, thereby establishing Abia as a critical sentinel for sub-Saharan Africa’s resistance pattern. By integrating longitudinal molecular trends with clinical outcomes and predictive modelling, and forecasting a 15% surge in failures by 2030, these findings inform Nigeria’s Malaria Strategic Plan and advocate timely shifts to triple ACTs that could avert thousands of cases annually at cost-effective scales, while pioneering scalable genomic tools for resource-limited settings to fortify global malaria elimination efforts against evolving drug pressures.
ACKNOWLEDGMENT
We express our gratitude to the study participants in Abia State, Nigeria, whose invaluable contributions made this research possible, and to the tertiary health facilities for their unwavering support in sample collection and clinical assessments, to the Abia State Ministry of Health Ethics Committee for ethical oversight and to local community leaders for facilitating engagement. Finally, we acknowledge the tireless efforts of our field enumerators and laboratory technicians, whose commitment advanced malaria surveillance in this region.
REFERENCES
- Ochola, R., 2025. The case for genomic surveillance in Africa. Trop. Med. Infect. Dis., 10.
- Aliyu, I.A., J.A. Bala, B.M. Musa, T.G. Amole and A.B. Ahmed et al., 2025. Molecular epidemiology of Plasmodium falciparum drug resistance and vaccine targets in high-transmission settings in Africa. Sci. Rep., 16.
- Amusan, A., O. Akinola, K. Akano, M. Hernández-Castañeda and J.K. Dick et al., 2025. Polymorphisms in Pfkelch13 domains before and after the introduction of artemisinin-based combination therapy in Southwest Nigeria. PLoS ONE, 20.
- Bamikole, O.J., A.F. Fayehun, Y.A. Uthman, I.A. Salako, B.A. Adedeji, M.D.B. Olufeagba and O.K. Amodu, 2025. Molecular surveillance of antimalarial drug resistance genes in Nigeria: A systematic review and roadmap to malaria elimination. BMC Med. Genomics, 18.
- Enebeli, U.U., A.N. Amadi, O.K. Iro, E.T. Oparaocha and E.A. Nwoke et al., 2019. Assessment of water, sanitation and hygiene practices and the occurrence of childhood malaria in Abia State, Nigeria. Researchjournali’s J. Public Health, 5.
- Ozodiegwu, I.D., M. Ambrose, B. Galatas, M. Runge and A. Nandi et al., 2023. Application of mathematical modelling to inform national malaria intervention planning in Nigeria. Malar. J., 22.
- Guissou, R.M., J.M.T. Kaboré, I. Soulama, D. Hien and Y. Nombré et al., 2025. Multiple first-line therapeutic strategies to mitigate artemisinin resistance: Cost analysis of a pilot study from a health system perspective in Kaya health district, Burkina Faso. Malar J., 24.
- Ikegbunam, M., V. Tzirtziganis, M. Rodi, L. Anagu, L.P. de Carvalho, J. Inoue and J. Held, 2025. No evidence of molecular markers of piperaquine resistance in Southeastern Nigeria. Malar J., 24.
- MalariaGEN, M.M.A. Abdel Hamid, M.H. Abdelraheem, D.O. Acheampong and I. Adam et al., 2025. Pf8: An open dataset of Plasmodium falciparum genome variation in 33,325 worldwide samples. Wellcome Open Res., 10.
- Ramírez, A.M., A.A. Akindele, V.G. Mora, L. García and N. Lara et al., 2025. Mutational profile of pfdhfr, pfdhps, pfmdr1, pfcrt and pfk13 genes of P. falciparum associated with resistance to different antimalarial drugs in Osun State, Southwestern Nigeria. Trop. Med. Health, 53.
- Ogbonna, D.N., T.G. Sokari and A.A. Agomuoh, 2008. Antimalarial activities of some selected traditional herbs from South Eastern Nigeria against Plasmodium species. Res. J. Parasitol., 3: 25-31.
- Olasehinde, G.I., D.O. Ojurongbe, O.J. Akinjogunla, L.O. Egwari and A.O. Adeyeba, 2015. Prevalence of malaria and predisposing factors to antimalarial drug resistance in Southwestern Nigeria. Res. J. Parasitol., 10: 92-101.
- Oyebola, K.M., F.C. Ligali, A.J. Owoloye, O.O. Aina and Y.M. Alo et al., 2024. Assessing the therapeutic efficacy of artemether-lumefantrine for uncomplicated malaria in Lagos, Nigeria: A comprehensive study on treatment response and resistance markers. Malar. J., 23.
- Oyegbade, S.A., E.O. Mameh, D.O. Balogun, V.G.O. Aririguzoh and P.A. Akinduti, 2025. Emerging Plasmodium falciparum K13 gene mutation to artemisinin-based combination therapies and partner drugs among malaria-infected population in Sub-Saharan Africa. Parasites Hosts Dis., 63: 109-122.
- Umar, R.A., S.W. Hassan, M.J. Ladan, M.N. Jiya and I.K. Matazu et al., 2008. Therapeutic efficacy of chloroquine for uncomplicated Plasmodium falciparum malaria in Nigerian children at the time of transition to artemisinin-based combination therapy. Res. J. Parasitol., 3: 32-39.
- WHO, 2007. Methods and Techniques for Clinical Trials on Antimalarial Drug Efficacy: Genotyping to Identify Parasite Populations. World Health Organization, Amsterdam, The Netherlands, ISBN: 978 92 4 159630 5, Pages: 54.
- WHO, 2021. Global Technical Strategy for Malaria 2016-2030. World Health Organization, Switzerland, ISBN: 978 92 4 003135 7, Pages: 40.
- WHO, 2024. World Malaria Report 2024: Addressing Inequity in the Global Malaria Response. World Health Organization, Geneva, Switzerland, ISBN-13: 978-92-4-010444-0, Pages: 316.
- WHO, 2025. World Malaria Report 2025: Addressing the Threat of Antimalarial Drug Resistance. World Health Organization, Geneva, Switzerland, ISBN: 978-92-4-011782-2, Pages: 213.
- WHO, 2025. Compendium of Molecular Markers for Antimalarial Drug Resistance. World Health Organization, Geneva, Switzerland, Pages: 7.
How to Cite this paper?
APA-7 Style
Enebeli,
U.U., Kalu,
E.I., Amadi,
A.N., Kalu,
F.A., Igwe,
P.C., Kalu,
J.J., Kalu,
B.O. (2026). Genomic Surveillance of Plasmodium falciparum in Southeastern Nigeria: Implications for Artemisinin-Based Therapy Policy. Research Journal of Parasitology, 21(1), 1-8. https://doi.org/10.3923/parasite.2026.01.08
ACS Style
Enebeli,
U.U.; Kalu,
E.I.; Amadi,
A.N.; Kalu,
F.A.; Igwe,
P.C.; Kalu,
J.J.; Kalu,
B.O. Genomic Surveillance of Plasmodium falciparum in Southeastern Nigeria: Implications for Artemisinin-Based Therapy Policy. Res. J. Parasitol 2026, 21, 1-8. https://doi.org/10.3923/parasite.2026.01.08
AMA Style
Enebeli
UU, Kalu
EI, Amadi
AN, Kalu
FA, Igwe
PC, Kalu
JJ, Kalu
BO. Genomic Surveillance of Plasmodium falciparum in Southeastern Nigeria: Implications for Artemisinin-Based Therapy Policy. Research Journal of Parasitology. 2026; 21(1): 1-8. https://doi.org/10.3923/parasite.2026.01.08
Chicago/Turabian Style
Enebeli, Ugo, Uwadiako, Eziyi Iche Kalu, Agwu Nkwa Amadi, Faith Adamma Kalu, Perfection Chinyere Igwe, Justin Junior Kalu, and Beauty Olamma Kalu.
2026. "Genomic Surveillance of Plasmodium falciparum in Southeastern Nigeria: Implications for Artemisinin-Based Therapy Policy" Research Journal of Parasitology 21, no. 1: 1-8. https://doi.org/10.3923/parasite.2026.01.08

This work is licensed under a Creative Commons Attribution 4.0 International License.


